1.27 Million Deaths a Year: Why Antimicrobial Resistance Is a Global Emergency
Antimicrobial resistance (AMR) is no longer a distant threat on the medical horizon. It is a crisis unfolding right now, killing more people each year than HIV/AIDS or malaria. According to a landmark systematic analysis published in The Lancet, bacterial AMR directly caused 1.27 million deaths globally in 2019 and contributed to nearly 4.95 million total deaths that same year.
The trajectory is worsening. Projections from the same analysis estimate that between 2025 and 2050, 39 million deaths will be directly attributable to bacterial AMR, with associated deaths reaching 169 million. That translates to roughly three lives lost every minute for the next quarter century.
Key Fact: Antibiotic resistance rose in more than 40% of bacteria-drug combinations tracked between 2018 and 2023, with average annual increases of 5-15%, according to the WHO's Global Antibiotic Resistance Surveillance System.
The U.S. Centers for Disease Control and Prevention (CDC) reports that over 2.8 million antimicrobial-resistant infections occur annually in the United States alone, causing more than 35,000 deaths. The COVID-19 pandemic reversed a decade of progress: six major hospital-onset resistant bacteria increased 20% during the pandemic compared to pre-pandemic levels.
| AMR Statistic | Value | Source |
|---|---|---|
| Direct AMR deaths globally (2019) | 1.27 million | The Lancet |
| AMR-associated deaths globally (2019) | 4.95 million | The Lancet |
| Projected direct AMR deaths (2025-2050) | 39 million | The Lancet |
| U.S. resistant infections annually | 2.8+ million | CDC |
| U.S. AMR-related deaths annually | 35,000+ | CDC |
| NDM-CRE surge in U.S. (2019-2023) | 460% increase | CDC |
| Annual U.S. healthcare cost of AMR | $4.6+ billion | CDC |
Perhaps most alarming is the antibiotic pipeline crisis. The World Health Organization (WHO) reports that only 90 antibiotics are currently in clinical development, just 15 qualify as innovative, and only one candidate is in Phase III trials targeting WHO's critical-priority pathogens. With traditional drug development stalling, researchers are turning to radically different approaches — including genetic engineering tools that can neutralize resistance at its source.
The Hidden Highway: How Bacteria Share Resistance Genes Through Plasmids
To understand why AMR is so difficult to control, you need to understand how bacteria acquire and share resistance in the first place. Unlike humans, bacteria do not need to reproduce to pass along genetic traits. They can exchange DNA directly through a process called horizontal gene transfer.
The primary vehicles for this exchange are plasmids — small, circular DNA molecules that exist separately from a bacterium's main chromosome. Plasmids frequently carry antibiotic resistance genes, and they can replicate independently within a bacterial cell. When a bacterium divides, copies of these plasmids are passed to daughter cells. But plasmids can also jump between entirely different species of bacteria through three main mechanisms:
- Conjugation: A donor bacterium physically connects to a recipient through a tube-like structure called a pilus and directly transfers a copy of the plasmid. This is the most common route for spreading multi-drug resistance.
- Transformation: Bacteria absorb free-floating DNA fragments from their environment, including plasmid DNA released by dead cells.
- Transduction: Bacteriophages (viruses that infect bacteria) accidentally package plasmid DNA and inject it into new hosts.
This means a single resistant bacterium in a hospital, a farm, or even the human gut can transmit resistance to entirely different bacterial species within hours. The beneficial bacteria in your gut microbiome can receive resistance plasmids from pathogenic bacteria — and vice versa. Resistance does not stay contained; it flows through microbial communities like information through a network.
The clinical consequences are severe. Carbapenem-resistant Enterobacterales (CRE), which carry resistance on plasmids encoding genes like blaKPC and blaNDM, have mortality rates exceeding 40% in bloodstream infections. These plasmids often carry resistance to multiple antibiotic classes simultaneously, leaving physicians with few or no treatment options.
Quick Fact: A single resistance plasmid can carry genes conferring resistance to five or more antibiotic classes, turning a treatable infection into a potentially fatal one overnight.
Can CRISPR-Cas Technology Outsmart Drug-Resistant Superbugs?
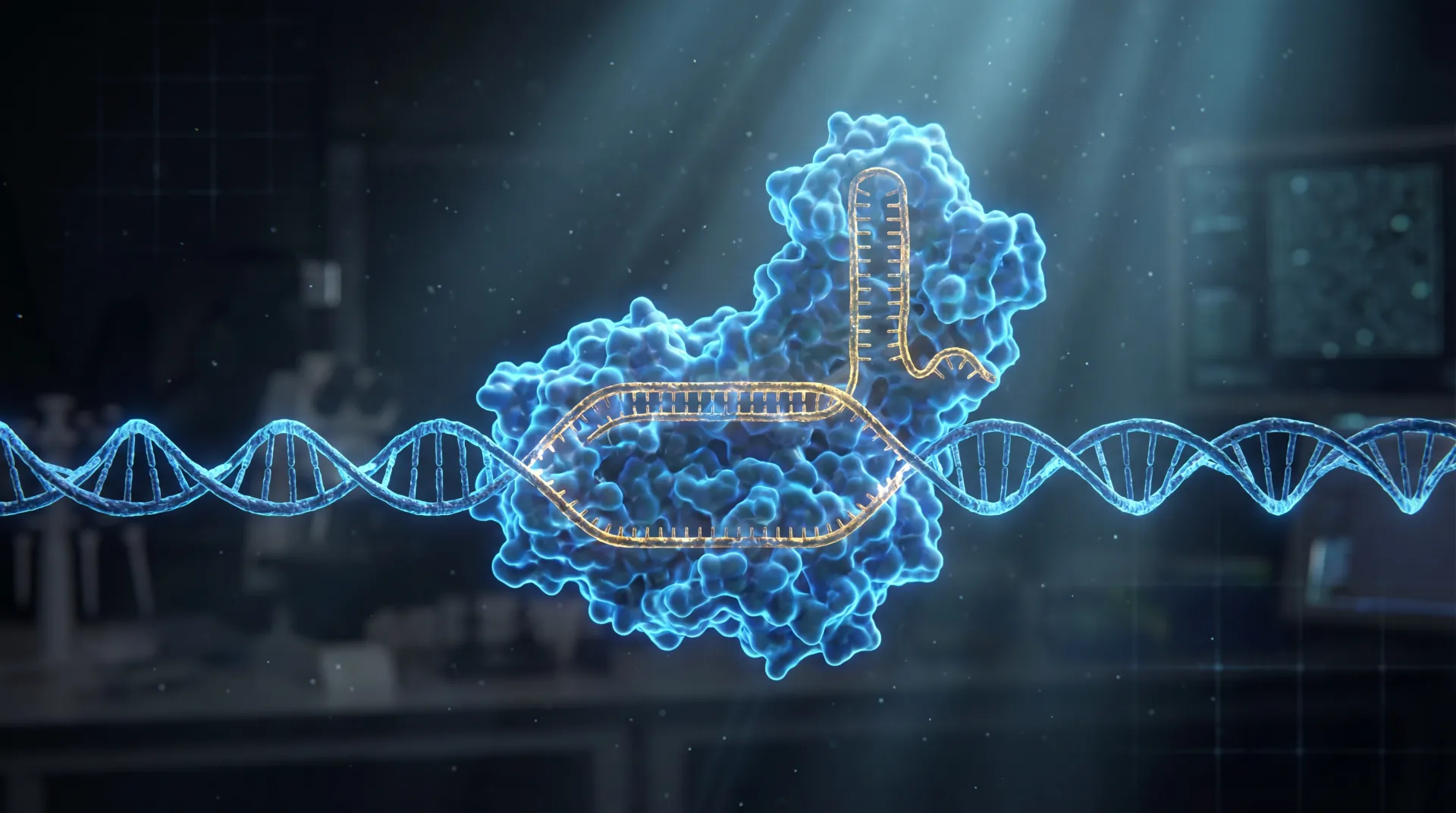
CRISPR-Cas (Clustered Regularly Interspaced Short Palindromic Repeats) systems are best known for their revolutionary applications in human gene therapy, but researchers have increasingly turned these molecular scissors toward a different target: the resistance genes that make bacteria so dangerous.
Unlike traditional antibiotics, which work by broadly disrupting bacterial cell processes and inevitably creating selection pressure for resistance, CRISPR-Cas systems can be programmed to target specific DNA sequences with extraordinary precision. When applied to AMR, this precision enables two distinct antimicrobial strategies.
Direct killing involves programming CRISPR-Cas to cut essential chromosomal genes in specific pathogenic bacteria, creating lethal DNA breaks that the targeted organism cannot repair. This selectively eliminates resistant strains while preserving beneficial bacteria — a major advantage over broad-spectrum antibiotics that can devastate the diverse probiotic strains that protect human health.
Plasmid curing takes a more elegant approach. Instead of killing the bacterium, CRISPR-Cas targets and destroys the resistance genes carried on plasmids. When the plasmid DNA is cut at the resistance gene, the plasmid is either degraded entirely or the gene is rendered nonfunctional. The bacterium survives but loses its resistance, becoming susceptible to conventional antibiotics again. Research has demonstrated curing efficiencies exceeding 94% for carbapenem resistance plasmids using CRISPR-Cas9.
| CRISPR System | Type | Mechanism | Primary Use in AMR |
|---|---|---|---|
| Cas9 | Type II | Blunt-end double-strand DNA break | Plasmid curing, gene inactivation |
| Cas12a | Type V | Staggered double-strand DNA break | Broader PAM compatibility for diverse targets |
| Cas3 | Type I | Progressive DNA degradation | Complete plasmid destruction |
| Cas13 | Type VI | RNA targeting | Diagnostics, transcript degradation |
The critical challenge has been delivery. CRISPR components must physically enter the target bacterium to work. Researchers have explored nanoparticles, engineered bacteriophages, and conjugative plasmids as delivery vehicles — each with distinct trade-offs between efficiency, host range, and scalability. This is precisely the challenge that the pPro-MobV system was designed to solve.
Inside pPro-MobV: The Gene Drive That Hunts Antibiotic Resistance
In early 2026, researchers at UC San Diego published a breakthrough that could fundamentally change how we fight antimicrobial resistance. The team, led by professors Ethan Bier and Justin Meyer, developed pPro-MobV — a conjugative CRISPR-based system that functions like a gene drive in bacteria, autonomously spreading through bacterial populations to seek out and destroy antibiotic resistance genes.
The study, published in npj Antimicrobials and Resistance, builds on the team's earlier Pro-AG (Pro-Active Genetics) system, which was first described in a 2019 Nature Communications paper. That original system demonstrated a self-amplifying genetic cassette capable of achieving a 100,000-fold reduction in antibiotic-resistant bacteria. The new pPro-MobV system adds something the original lacked: the ability to physically move between bacterial cells on its own.
The pPro-MobV plasmid, approximately 65 kilobases in size, integrates four key components into a single mobile genetic element:
- An inducible lambda-Red-Cas9 operon — combining the DNA-cutting power of Cas9 with phage-derived recombination proteins that enable precise genetic insertion
- A constitutive sgRNA (single guide RNA) — programmed to target a specific resistance gene (in this case, the bla gene conferring ampicillin resistance)
- Conjugation machinery from the IncP RK2 system — providing the molecular apparatus for cell-to-cell transfer across a broad range of Gram-negative bacteria
- An oriT (origin of transfer) site — enabling the plasmid to be mobilized during conjugation
The system operates in a cascading sequence. First, the pPro-MobV plasmid transfers from a donor cell to a target bacterium through conjugation — essentially bacterial mating. Once inside, Cas9 locates and cuts the resistance gene on the target plasmid. The lambda-Red recombination system then inserts the Pro-AG cassette directly into the cut site, permanently inactivating the resistance gene. This insertion creates a self-amplifying feedback loop: the inserted cassette becomes a template for further copying, boosting editing efficiency two orders of magnitude higher than simpler cut-and-destroy approaches.
The results were striking. The pPro-MobV system reduced antibiotic resistance prevalence by three to five logs (a 1,000 to 100,000-fold reduction), depending on the bacterial strain tested. The system also proved effective inside biofilms — dense bacterial communities that are notoriously resistant to antibiotics and disinfectants.
In the Researchers' Words: "With this new CRISPR-based technology we can take a few cells and let them go to neutralize antibiotic resistance in a large target population." — Ethan Bier, UC San Diego
The researchers also built in a safety mechanism called Homology-Based Deletion (HBD). This CRISPR-driven process can precisely remove the Pro-AG cassette using short direct repeats flanking the inserted DNA. HBD components can be delivered via a secondary plasmid or phage, functioning as a biological kill switch that prevents uncontrolled spread. Because Pro-AG and HBD are differentially regulated by the bacterial RecA pathway, they can be independently controlled — a critical feature for any self-propagating genetic system.
Antibiotics vs. CRISPR: Which Approach Wins the War on Resistance?
Traditional antibiotics and CRISPR-based approaches represent fundamentally different philosophies for fighting bacterial infections. Understanding their differences reveals why genetic engineering may be essential to complementing — though not replacing — conventional treatments.
Broad-spectrum antibiotics act as chemical weapons, disrupting essential bacterial processes like cell wall synthesis or protein production. They work quickly and have saved hundreds of millions of lives. But their indiscriminate nature creates two serious problems. First, they destroy beneficial bacteria alongside pathogens, disrupting the immune-supporting microbiome and opening the door to secondary infections like Clostridioides difficile. Second, they create intense selection pressure: any bacterium that survives an antibiotic course — thanks to a resistance gene on a plasmid — reproduces to fill the ecological vacuum left by dead competitors.
| Feature | Traditional Antibiotics | pPro-MobV / CRISPR-Cas |
|---|---|---|
| Specificity | Broad-spectrum; damages beneficial bacteria | Programmable; targets only specific resistance genes |
| Microbiome impact | Significant disruption; secondary infection risk | Preserves beneficial microbiota |
| Resistance evolution | Creates selection pressure favoring resistance | Actively reverses resistance at the genetic level |
| Biofilm effectiveness | Poor penetration (100-1000x higher tolerance) | Demonstrated effectiveness inside biofilms |
| Population-level action | Requires treating each individual patient | Self-propagating through bacterial populations |
| Re-sensitization ability | Cannot restore antibiotic susceptibility | Cures resistance plasmids, restoring sensitivity |
| Development pipeline | Only 90 candidates in clinical trials | Programmable against any sequenced resistance gene |
| Reversibility | Ecological disruption difficult to undo | HBD safeguard enables removal of genetic cassette |
The CRISPR approach does not replace antibiotics. Instead, it addresses the fundamental weakness of antibiotic therapy: while antibiotics kill bacteria, they cannot undo the genetic basis of resistance. The pPro-MobV system can. As Justin Meyer has noted, this technology represents "one of the few ways that can actively reverse the spread of antibiotic-resistant genes." This capacity for re-sensitization — restoring bacteria to a state where existing antibiotics work again — could extend the useful life of our current antibiotic arsenal by decades.
For individuals already focused on strengthening their immune systems naturally, CRISPR-based approaches offer an additional layer of protection: targeted elimination of resistance genes without the collateral damage to gut health that accompanies every course of antibiotics.
What Stands Between the Laboratory and Your Medicine Cabinet?
Despite its remarkable promise, the pPro-MobV system faces significant hurdles before it could reach clinical use. The gap between laboratory proof-of-concept and approved medical therapy is vast, and several technical, biological, and regulatory challenges must be addressed.
Host range limitations represent a major technical barrier. The IncP RK2 conjugation system used by pPro-MobV transfers efficiently across Gram-negative bacteria, but it cannot penetrate Gram-positive organisms. This means pathogens like methicillin-resistant Staphylococcus aureus (MRSA) and vancomycin-resistant Enterococcus (VRE) — both critical WHO priority pathogens — remain beyond the system's current reach.
Bacterial counter-defenses are another concern. Bacteria have evolved their own anti-CRISPR mechanisms, including proteins that inhibit Cas9 activity, mutations in PAM (Protospacer Adjacent Motif) sequences that prevent CRISPR recognition, and DNA methylation that masks target sites. While resistance to CRISPR targeting occurs at frequencies around one in ten thousand, the scale of bacterial populations in clinical settings means such escape mutants could emerge.
Multiplexing challenges present a practical limitation. The current pPro-MobV system targets a single resistance gene (bla). Multi-drug resistant bacteria often carry resistance to five or more antibiotic classes across multiple plasmids and chromosomal insertions. Effective clinical deployment would require multiplexed sgRNAs targeting several genes simultaneously — significantly increasing the engineering complexity.
Regulatory uncharted territory may be the tallest barrier. No regulatory framework currently exists for approving self-propagating genetic systems for use in human medicine or environmental applications. The FDA, EMA, and other agencies would need to develop entirely new evaluation criteria for a therapy that, by design, spreads autonomously and modifies the genomes of organisms in its path. The HBD safety mechanism addresses some biosafety concerns, but validation in complex real-world environments remains incomplete.
In vivo validation is still in early stages. While the system has shown impressive results in laboratory cultures and biofilm models, the leap to animal models — and eventually human clinical trials — requires demonstrating safety, efficacy, and predictability in the enormously complex environment of the human body. The diversity of bacterial pathogens in conditions like food poisoning illustrates just how varied the microbial communities are that this system would need to navigate.
Researchers have identified several promising next steps: engineering multiplexed sgRNA arrays for multi-target capability, combining conjugative delivery with bacteriophage vectors for broader penetration, and exploring applications in controlled environments like hospital surfaces and wastewater treatment before attempting direct clinical use.
Frequently Asked Questions
What is the pPro-MobV CRISPR-Cas system?
The pPro-MobV system is a genetic engineering tool developed at UC San Diego that combines CRISPR-Cas9 gene editing with conjugative transfer machinery. This allows it to spread autonomously through bacterial populations and destroy antibiotic resistance genes. It functions like a gene drive for bacteria — a small number of engineered cells can neutralize resistance across a large population, achieving 1,000 to 100,000-fold reductions in resistance prevalence.
How does CRISPR target antibiotic resistance without killing beneficial bacteria?
CRISPR-Cas systems are programmable to target specific DNA sequences. When aimed at a resistance gene on a plasmid, the system cuts and destroys or inactivates that gene without affecting the bacterium's essential functions or other bacteria in the community. This is fundamentally different from antibiotics, which broadly kill bacteria regardless of whether they carry resistance genes or play beneficial roles in the gut microbiome.
Could bacteria develop resistance to CRISPR-based treatments?
Yes, bacteria can potentially evade CRISPR targeting through mutations in the recognition sequence, expression of anti-CRISPR proteins, or DNA methylation. However, these escape mechanisms occur at relatively low frequencies (approximately 1 in 10,000), and the self-amplifying nature of the pPro-MobV system — combined with the ability to reprogram guide RNAs against new targets — makes it more adaptable than fixed-compound antibiotics.
When could CRISPR-based antimicrobial therapies become available?
Clinical availability is likely years to decades away. The pPro-MobV system has been validated in laboratory cultures and biofilm models but has not yet entered animal testing. Regulatory frameworks for self-propagating genetic therapies do not yet exist. Near-term applications may emerge first in controlled environments like hospital surfaces, water treatment facilities, or agricultural settings before direct clinical use in human patients.
Is antimicrobial resistance really as dangerous as cancer or heart disease?
By the numbers, AMR is already deadlier than many well-known diseases. The 1.27 million annual deaths directly caused by resistant bacteria exceed the death toll from HIV/AIDS (approximately 860,000) and malaria (approximately 620,000). Projections suggest AMR-attributable deaths will nearly double by 2050, and without intervention, resistant infections could cause $1-3.4 trillion in annual GDP losses globally.
Related Articles
- Health Benefits of Probiotics — How beneficial bacteria support digestion, immunity, and overall health.
- Probiotic Strains and Species: Benefits and Research — A detailed look at the specific bacterial strains that promote gut health.
- Immune-Boosting Probiotic Foods — Foods that help strengthen the immune system through beneficial microorganisms.
- Natural Ways to Strengthen Your Immune System During Winter — Evidence-based strategies for supporting immune function year-round.
- Food Poisoning: Causes, Symptoms, Treatments, and Recovery — Understanding foodborne pathogens and their treatment.
Medical Disclaimer
This article is for informational and educational purposes only and is not medical advice, diagnosis, or treatment. Always consult a licensed physician or qualified healthcare professional regarding any medical concerns. Never ignore professional medical advice or delay seeking care because of something you read on this site. If you think you have a medical emergency, call 911 immediately.